
The transcriptome pipelines run on the Wexac cluster. In order to run a new transcriptome analysis, your fastq files must be in your Collaboration folder on Wexac, in the correct structure (UTAP requirements & description) .
To run a new analysis from existing files in the Collaboration folder login to http://utap.wexac.weizmann.ac.il using your Weizmann userID and password and click on "Run pipeline":
This page concentrates on the common aspects of the RNA-Seq, MARS-Seq or SCRB-Seq pipelines.
1. For MARS-Seq, you will get this screen:

2. For RNA-seq, you will get this screen:

Select whether or not your protocol is stranded (the sequenced reads saves the original strand of RNA fragments) or non-stranded.
Specify your adapters for each read (R1 and R2). These adapters will be removed from the reads by the pipeline. The default adapters are the Illumina compatible TruSeq i5 and i7 adapters.
3. For SCRB-Seq, you will get this screen: - will be added
Fill in the project name, select the genome and annotation.
Browse within your Collaboration directory structure and Select the root folder of the sub-folders of the samples for analysis with the appropriate button.
Note that if you wish to go up one level (or more), please click the desired folder level on the path at the top of the window.
If there is an error with the folder you selected, you must fix it in order to run the pipeline.
Select the desired output folder.
Continue by filling in all of the fields.
C. Select DESeq2 options for the statistical analysis
Select "Run DESeq2" if you desire to identify differentially expressed genes using the DESeq2 package as described in the DESeq2 manual. If you select this option, al least two categories must be defined (by filling in the category names).
Associate each sample to a category, use the arrows to move the sample to the appropriate category.
You may add additional categories by using the relevant buttons.

If the samples were prepared in different batches, you can add this information: After moving the samples into categories boxes, click on the "Add Batch Effect" button, then select the samples that belong to one batch and click on the "Batch 1" button. Repeat the operation with the other groups of the samples.

All the steps of the pipeline (mapping, counts etc.) will be run on all of the samples, except for DESeq2 which will be run only on the samples with categories.
Finally, submit the run for analysis.
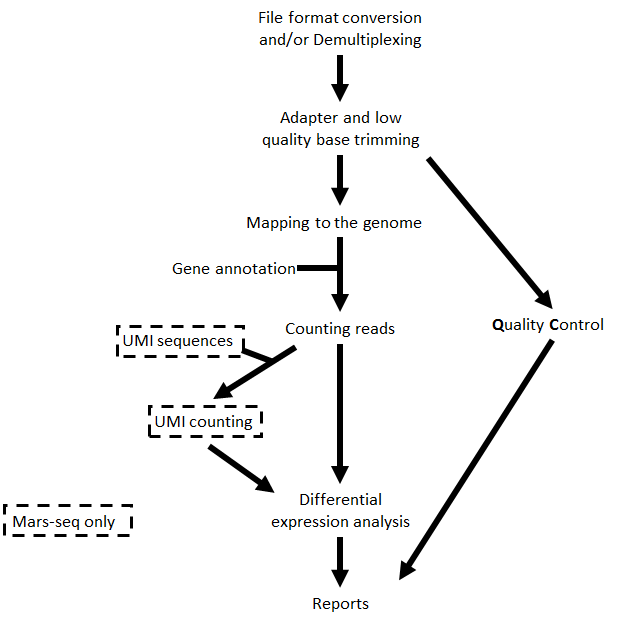
The steps performed by the pipeline include:
Steps 4 and 6 are performed only for MARS-Seq
Upon completion, you will get an email with links to the results report. For an interactive detailed explanation of the report use the relevant e-learning module.
The report includes several sections:
For the counts of the reads per gene we use with annotation files (gtf format) from RefSeq or GENCODE (more elaborate i.e. contains more genes and transcripts) . In MARS-Seq analysis we use a modified version of the gene that includes 1000 bp upstream of the TES (transcription end site) on the transcript and 100 bp downstream of the TES.
Transcriptome pipeline for Weizmann Institute users: http://utap.wexac.weizmann.ac.il
Kohen et al. BMC Bioinformatics (2019) 20:154 https://doi.org/10.1186/s12859-019-2728-2 (PMID: 30909881)